Mahotas – Haralick features
Last Updated :
16 May, 2022
In this article we will see how we can get the haralick features of image in mahotas. Haralick texture features are calculated from a Gray Level Co-occurrence Matrix, (GLCM), a matrix that counts the co-occurrence of neighboring gray levels in the image. The GLCM is a square matrix that has the dimension of the number of gray levels N in the region of interest (ROI). For this we are going to use the fluorescent microscopy image from a nuclear segmentation benchmark. We can get the image with the help of command given below
mahotas.demos.nuclear_image()
Below is the nuclear_image
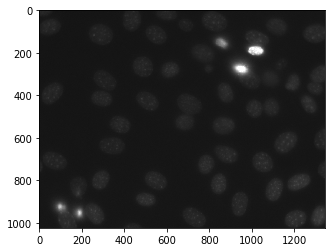
In order to do this we will use mahotas.features.haralick method
Syntax : mahotas.features.haralick(img)
Argument : It takes image object as argument
Return : It returns numpy.ndarray
Note : The input of the this should be the filtered image or loaded as grey
In order to filter the image we will take the image object which is numpy.ndarray and filter it with the help of indexing, below is the command to do this
image = image[:, :, 0]
Example 1 :
Python3
import mahotas
import mahotas.demos
import mahotas as mh
import numpy as np
from pylab import imshow, show
nuclear = mahotas.demos.nuclear_image()
nuclear = nuclear[:, :, 0]
nuclear = mahotas.gaussian_filter(nuclear, 4)
threshed = (nuclear > nuclear.mean())
labeled, n = mahotas.label(threshed)
print("Labelled Image")
imshow(labeled)
show()
h_feature = mahotas.features.haralick(labelled)
print("Haralick Features")
imshow(h_feature)
show()
|
Output :

Example 2 :
Python3
import numpy as np
import mahotas
from pylab import imshow, show
img = mahotas.imread('dog_image.png')
img = img[:, :, 0]
gaussian = mahotas.gaussian_filter(img, 15)
gaussian = (gaussian > gaussian.mean())
labeled, n = mahotas.label(gaussian)
print("Labelled Image")
imshow(labelled)
show()
h_feature = mahotas.features.haralick(labelled)
print("Haralick Features")
imshow(h_feature)
show()
|
Output :

Share your thoughts in the comments
Please Login to comment...